Our cells are crisscrossed by a system of membrane tubes and pockets called the endoplasmic reticulum (ER). It is crucial for the production of biomolecules and is continuously built up and degraded. Degradation, known as ER-phagy, is promoted by the protein ubiquitin, which controls many processes in the cell. If the proteins involved in ER-phagy are defective, neurodegenerative diseases result. This has been discovered by an international research team led by Goethe University Frankfurt (as part of the EMTHERA research cluster) and Jena University Hospital and published in two papers in the renowned journal Nature.
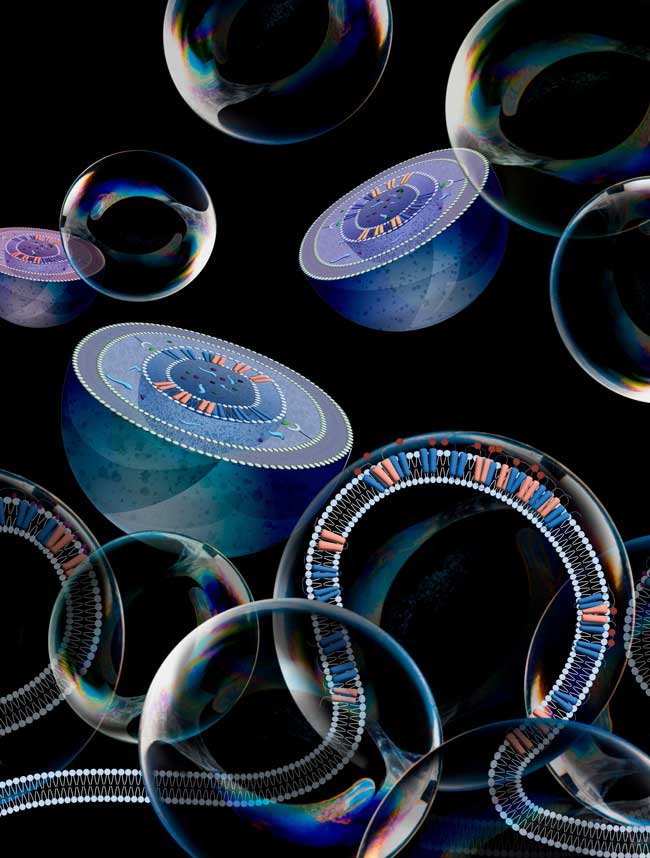

A tangle of pockets, tubes and sac-like membrane structures runs through the cells of humans, animals, plants and fungi: the endoplasmic reticulum, or ER for short. In the ER, proteins are manufactured, folded into their three-dimensional structure and modified, lipids and hormones are produced and calcium concentrations in the cell are controlled. In addition, the ER forms the basis for the cellular transport system, feeds misfolded proteins to intracellular disposal and renders toxins that have entered the cell harmless.
In view of its multiple tasks, the ER is constantly being remodeled. A process called ER-phagy (roughly “self-digestion of the ER”) is responsible for ER degradation. Involved is a group of signal-receiving proteins – receptors – that are responsible for the membrane curvatures of the ER and thus for its multiple forms in the cell. In ER-phagy, the receptors accumulate at specific sites on the ER and increase membrane curvature to such an extent that, as a consequence, part of the ER is strangulated and broken down into its component parts by cellular recycling structures (autophagosomes).
In cell culture experiments, biochemical and molecular biological studies, and by computer simulations, the scientific team led by Professor Ivan Đikić of Goethe University Frankfurt first tested the membrane curvature receptor FAM134B and demonstrated that ubiquitin promotes and stabilizes the formation of clusters of FAM134B protein in the ER membrane. Thus, ubiquitin drives ER-phagy. Đikić explains: “Ubiquitin causes the FAM134B clusters to become more stable and the ER to bulge out more at these sites. The stronger membrane curvature then leads to further stabilization of the clusters and, moreover, attracts additional membrane curvature proteins. So the effect of ubiquitin is self-reinforcing.” The researchers were also able to detect cluster formation using super-high resolution microscopy.
Đikić continues: “To fulfill this function, ubiquitin changes the shape of part of the FAM134B protein. This is another facet of ubiquitin that performs an almost unbelievable array of tasks to keep all different cell functions working.”
The importance of ER-phagy is demonstrated by diseases resulting from a defective FAM134B protein. A team led by Professor Christian Hübner from Jena University Hospital previously identified mutations in the FAM134B gene causing a very rare hereditary sensory and autonomic neuropathy (HSAN), in which sensory nerves die. As a result, patients are unable to perceive pain and temperature correctly, which can lead to incorrect stresses or injuries going unnoticed and developing into chronic wounds. In a long-standing collaboration between Jena University Hospital and Goethe University Frankfurt FAM134B was identified as the first receptor for ER-phagy.
Mutations in another membrane curvature protein called ARL6IP1 cause a similar neurodegenerative disorder which combines sensory defects with muscle hardening (spasticity) in the legs. The scientific team led by Christian Hübner and Ivan Đikić has now identified that ARL6IP1 belongs to the ER-phagy machinery as well and is also ubiquitinated during ER-phagy.
Christian Hübner explains: “In mice that do not possess the ARL6IP1 protein, we can see that the ER virtually expands and degenerates as the cells age. This leads to an accumulation of misfolded proteins or protein clumps, which are no longer disposed of in the cell. As a result, nerve cells in particular, which do not renew as quickly as other body cells, die, causing the clinical symptoms in affected patients and genetically modified mice. We hypothesize from our data that the two membrane curvature receptors FAM134B and ARL6IP1 form mixed clusters during ER-phagy and depend on each other to control normal size and function of ER. Additional work will be required to fully acknowledge the role of ER-phagy in neurons as well as in other cell types.”
Overall, however, the research teams have taken a decisive step toward understanding ER-phagy, Đikić is convinced: “We now understand better how cells control their functions and thus create something we call cellular homeostasis. In biology, this knowledge allows fascinating insights into the incredible achievements of our cells, and for medicine it is essential for understanding diseases, diagnosing them on time and helping patients by developing new therapies. ”
Scientists from the following institutions were involved in the work:
- Goethe University Frankfurt
- Jena University Hospital
- Max Planck Institute for Biophysics, Frankfurt
- Max Planck Institute of Biochemistry, Martinsried
- Fraunhofer Institute for Translational Medicine and Pharmacology, Frankfurt, Germany
- University Hospital RWTH Aachen
- Howard Hughes Medical Institute, San Diego, La Jolla, Calif.
- University of Groningen
- Aarhus University
Publications:
1) Alexis González, Adriana Covarrubias-Pinto, Ramachandra M. Bhaskara, Marius Glogger, Santosh K. Kuncha, Audrey Xavier, Eric Seemann, Mohit Misra, Marina E. Hoffmann, Bastian Bräuning, Ashwin Balakrishnan, Britta Qualmann, Volker Dötsch, Brenda A. Schulman, Michael M. Kessels, Christian A. Hübner, Mike Heilemann, Gerhard Hummer, Ivan Dikic: Ubiquitination regulates ER-phagy and remodeling of endoplasmic reticulum. Nature (2023) https://doi.org/10.1038/s41586-023-06089-2
2) Hector Foronda, Yangxue Fu, Adriana Covarrubias-Pinto, Hartmut T. Bocker, Alexis González, Eric Seemann, Patricia Franzka, Andrea Bock, Ramachandra M. Bhaskara, Lutz Liebmann, Marina E. Hoffmann, Istvan Katona, Nicole Koch, Joachim Weis, Ingo Kurth, Joseph G. Gleeson, Fulvio Reggiori, Gerhard Hummer, Michael M. Kessels, Britta Qualmann, Muriel Mari, Ivan Dikić, Christian A. Hübner: Heteromeric 1 clusters of ubiquitinated ER-shaping proteins drive ER-phagy. Nature (2023) https://doi.org/10.1038/s41586-023-06090-9